Materials:
- Chromebook
- pencil/pen
- internet
- blast webpage
- 4 different genes
Steps!
1. Go to the BLAST homepage: http://blast.ncbi.nlm.nih.gov/Blast.cgi
2. Click on “Saved Strategies” on the top of the page. Procedures:
3. Under “Upload Search Strategy,” click on “Browse” and locate one of the gene fi les you saved onto your computer.
4. Click “View.”
5. A screen will appear with the parameters for your query already confi gured.NOTE: Do not alter any of the parameters. Scroll down the page and click on the “BLAST” button at the bottom.6. After collecting and ana-lyzing all of the data for that particular gene (see instructions below), repeat this procedure for the other three gene sequences.
Step 3: Instructions for Analyzing BLAST Queries1. Scroll down to the section titled, “Sequences produc-
ing signifi cant alignments”. The list of organisms that appear below this section are those with sequences identical to or most similar to the gene of interest. The most similar sequences are listed fi rst and as you move down the list, the sequences become less similar to your gene of interest. 2. Try clicking on a particular species listed to get a full
report that includes the species’ classifi cation scheme, the research journal the gene was fi rst reported in, and the sequence of bases that appear to align with your gene of interest.
3. Click “Distance tree of results,” a cladogram of the species with similar sequences to your gene of interest placed on the cladogram will be shown according to how closely their matched gene aligns with your gene of interest.
2. Click on “Saved Strategies” on the top of the page. Procedures:
3. Under “Upload Search Strategy,” click on “Browse” and locate one of the gene fi les you saved onto your computer.
4. Click “View.”
5. A screen will appear with the parameters for your query already confi gured.NOTE: Do not alter any of the parameters. Scroll down the page and click on the “BLAST” button at the bottom.6. After collecting and ana-lyzing all of the data for that particular gene (see instructions below), repeat this procedure for the other three gene sequences.
Step 3: Instructions for Analyzing BLAST Queries1. Scroll down to the section titled, “Sequences produc-
ing signifi cant alignments”. The list of organisms that appear below this section are those with sequences identical to or most similar to the gene of interest. The most similar sequences are listed fi rst and as you move down the list, the sequences become less similar to your gene of interest. 2. Try clicking on a particular species listed to get a full
report that includes the species’ classifi cation scheme, the research journal the gene was fi rst reported in, and the sequence of bases that appear to align with your gene of interest.
3. Click “Distance tree of results,” a cladogram of the species with similar sequences to your gene of interest placed on the cladogram will be shown according to how closely their matched gene aligns with your gene of interest.
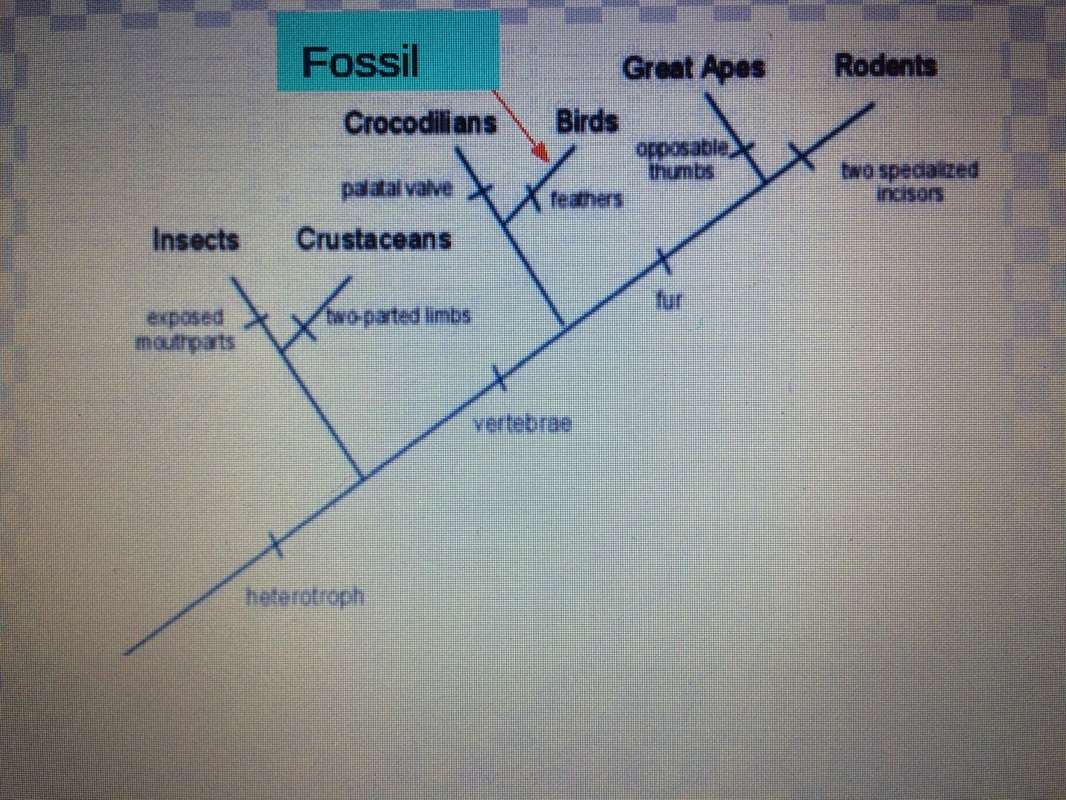
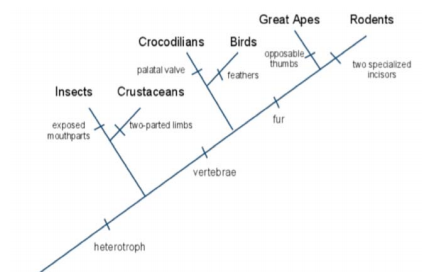
| Justification: Gene #1 | Based on the data I reaseached I found that gene #1 has the most similar gene sequences as Gallus, Gallus Collogagen, type V, alpha 1 (COL5A1) MRNA. This species is located under gene 1. The species and the fossil are 100% similar in gene sequences. Also I found that the species with less common gene sequences is Anolis Corolinesis. Then what I last identified was before the fossil and birds branches off they were first placentals. In the cladogram the fossil is seperated from birds and sea creatures such as sea turtles. |
Gene #2

Justification: | Based on my researched I found that the specie with the most similar gene sequence is the DRosophila Melanogaster. There are six species about gene 2. Gene similarity is 100% similar. Pediculus is the specie least similar to the fossil. In the cladogram it branches off from mosquitos and ants. Then goes to flies and unknown fossil. |
Gene #3
JUstification: | Based on my information I came across the specie with the most similar gene sequence as the fossil is Taeniopygia guttata. The specie is located where it branches off of bats and birds. The gene similarity is 95% right above the Taeniopygia Guttata. It has a gene similarity with bats and birds. The specie that is least similar because of gene sequence is an alligator. The cladogram goes from vertabraes, then breaks off to turtles and birds --> then Guttata and the unknown fossil. The fossil seperates from rabbits and hairs so it does not contain fur. |
Gene #4
Justification: | The specie who has the most common gene sequence is the alligator. The specie is located right under gene #4 in the cladogram. Between gene #4 and the fossil it is 100% similar in gene sequence. On the other side, the specie least related to this fossil is Barbourula busangensis. The cladogram break off from birds and vertabraes and unknown fossil is alligned with aligators. The fossil seperates from whales and dolphins so it is not a mammal. |










 RSS Feed
RSS Feed
